Info
tenx_
10x Genomics (2023)
246.08 MiB
23-09-2024
6434 cells × 19288 genes
Gene expression library of Mouse Embryo (CytAssist FFPE) using the Mouse Whole Transcriptome Probe Set
tenx_
10x Genomics (2023)
246.08 MiB
23-09-2024
6434 cells × 19288 genes
CREATED
23-09-2024
DIMENSIONS
6434 × 19288
The tissue was sectioned as described in Visium CytAssist Spatial Gene Expression for FFPE Tissue Preparation Guide Demonstrated Protocol CG000518. Tissue sections of 5 µm was placed on a standard glass slide, and H&E-stained following deparaffinization. Sections were coverslipped with 85% glycerol, imaged, decoverslipped, followed by dehydration & decrosslinking (Demonstrated Protocol CG000520). The glass slide with the tissue section was processed with the Visium CytAssist instrument to transfer analytes to a Visium CytAssist Spatial Gene Expression slide (11 mm Capture Area). The probe extension and library construction steps follow the standard Visium for FFPE workflow outside of the instrument.
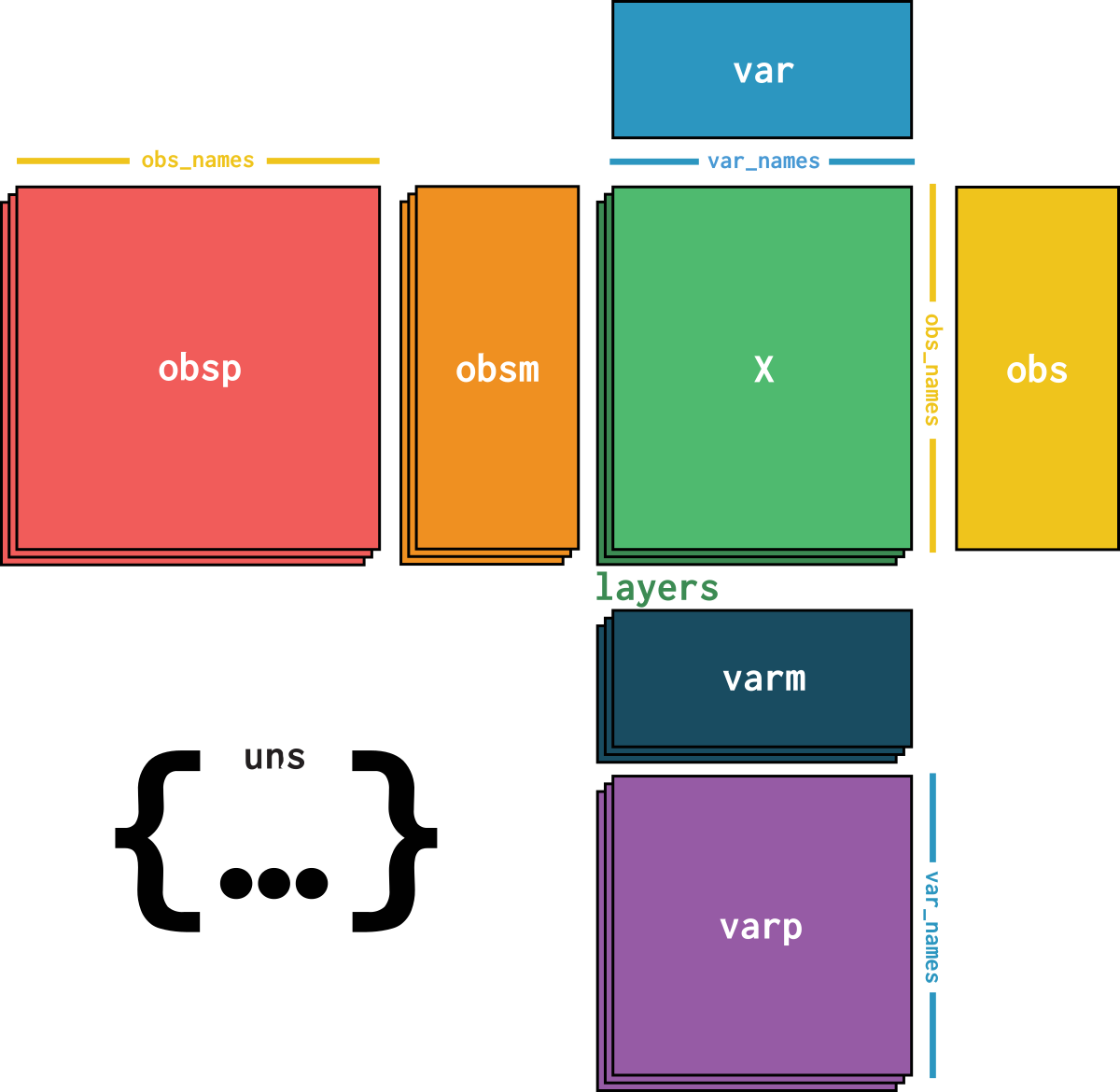
dataset is an AnnData object with n_obs × n_vars = 6434 × 19288 with slots:
feature_id, feature_namecountsdataset_description, dataset_id, dataset_name, dataset_organism, dataset_reference, dataset_summary, dataset_url| Name | Description | Type | Data type | Size |
|---|---|---|---|---|
| var | ||||
feature_
|
Unique identifier for the feature, usually a ENSEMBL gene id. |
vector
|
object
|
19288 |
feature_
|
A human-readable name for the feature, usually a gene symbol. |
vector
|
object
|
19288 |
| layers | ||||
counts
|
Raw counts |
sparsematrix
|
float32
|
6434 × 19288 |
| uns | ||||
dataset_
|
Long description of the dataset. |
atomic
|
str
|
1 |
dataset_
|
A unique identifier for the dataset. This is different from the obs.dataset_id field, which is the identifier for the dataset from which the cell data is derived.
|
atomic
|
str
|
1 |
dataset_
|
A human-readable name for the dataset. |
atomic
|
str
|
1 |
dataset_
|
The organism of the sample in the dataset. |
atomic
|
str
|
1 |
dataset_
|
Bibtex reference of the paper in which the dataset was published. |
atomic
|
str
|
1 |
dataset_
|
Short description of the dataset. |
atomic
|
str
|
1 |
dataset_
|
Link to the original source of the dataset. |
atomic
|
str
|
1 |
dataset.layers['counts']In R: dataset$layers[["counts"]]
Type: sparsematrix, data type: float32, shape: 6434 × 19288
Raw counts
dataset.uns['dataset_description']In R: dataset$uns[["dataset_description"]]
Type: atomic, data type: str, shape: 1
Long description of the dataset.
dataset.uns['dataset_id']In R: dataset$uns[["dataset_id"]]
Type: atomic, data type: str, shape: 1
A unique identifier for the dataset. This is different from the obs.dataset_id field, which is the identifier for the dataset from which the cell data is derived.
dataset.uns['dataset_name']In R: dataset$uns[["dataset_name"]]
Type: atomic, data type: str, shape: 1
A human-readable name for the dataset.
dataset.uns['dataset_organism']In R: dataset$uns[["dataset_organism"]]
Type: atomic, data type: str, shape: 1
The organism of the sample in the dataset.
dataset.uns['dataset_reference']In R: dataset$uns[["dataset_reference"]]
Type: atomic, data type: str, shape: 1
Bibtex reference of the paper in which the dataset was published.
dataset.uns['dataset_summary']In R: dataset$uns[["dataset_summary"]]
Type: atomic, data type: str, shape: 1
Short description of the dataset.
dataset.uns['dataset_url']In R: dataset$uns[["dataset_url"]]
Type: atomic, data type: str, shape: 1
Link to the original source of the dataset.
dataset.var['feature_id']In R: dataset$var[["feature_id"]]
Type: vector, data type: object, shape: 19288
Unique identifier for the feature, usually a ENSEMBL gene id.
dataset.var['feature_name']In R: dataset$var[["feature_name"]]
Type: vector, data type: object, shape: 19288
A human-readable name for the feature, usually a gene symbol.