Info
zenodo_
Kuppe et al. (2022)
24.46 MiB
23-09-2024
4518 cells × 10687 genes
Gene expression library of human heart using 10x Visium.
zenodo_
Kuppe et al. (2022)
24.46 MiB
23-09-2024
4518 cells × 10687 genes
DATASET ID
zenodo_spatial/visium/human_heart_myocardial_infarction_2
REFERENCE
Kuppe et al. (2022)
SIZE
24.46 MiB
CREATED
23-09-2024
DIMENSIONS
4518 × 10687
Frozen heart samples were embedded in OCT (Tissue-Tek) and cryosectioned (Thermo Cryostar). The 10-µm section was placed on the pre-chilled Optimization slides (Visium, 10X Genomics, PN-1000193) and the optimal lysis time was determined. The tissues were treated as recommended by 10X Genomics and the optimization procedure showed an optimal permeabilization time of 12 or 18 min of digestion and release of RNA from the tissue slide. Spatial gene expression slides (Visium, 10X Genomics, PN-1000187) were used for spatial transcriptomics following the Visium User Guides
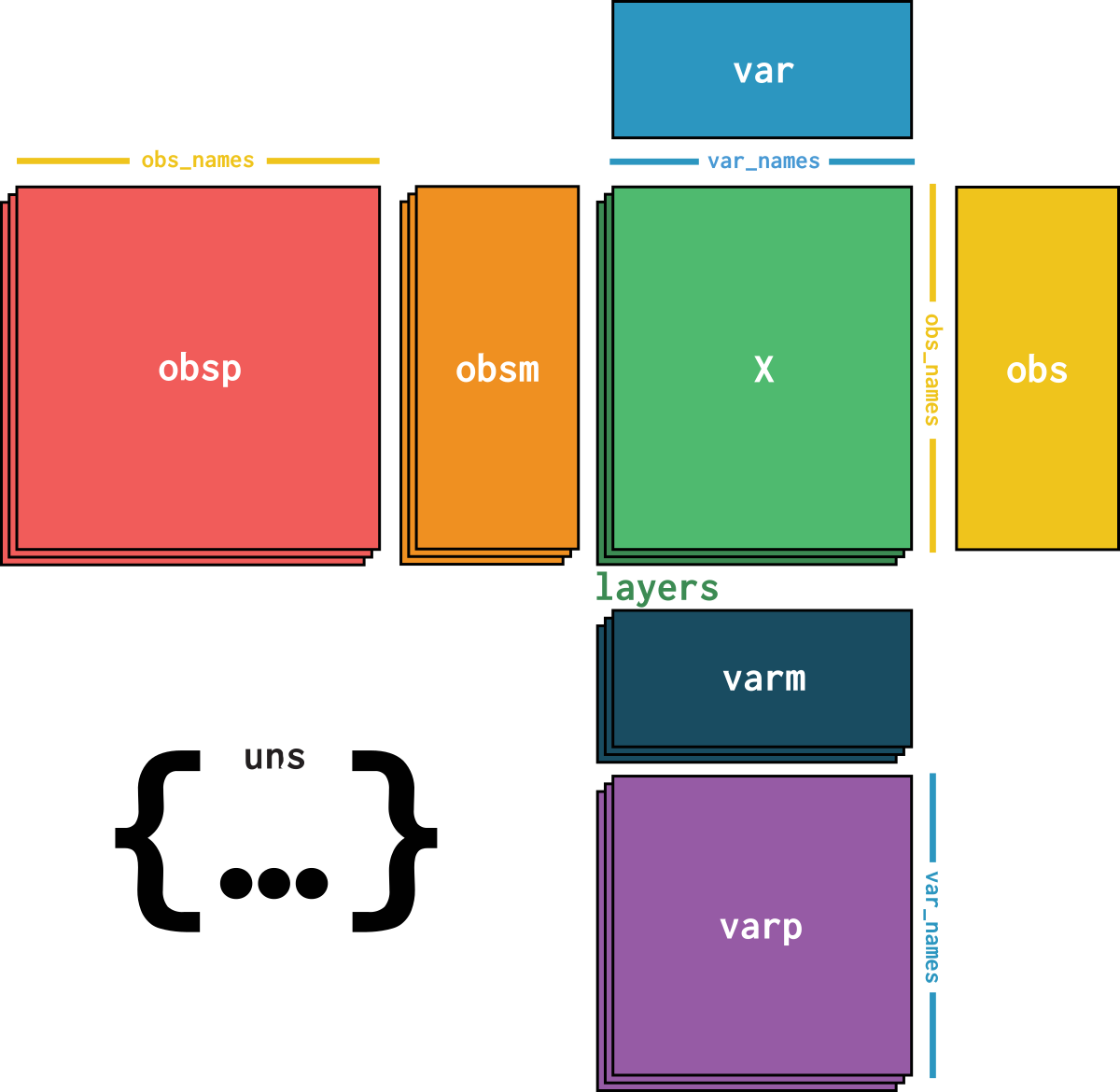
dataset is an AnnData object with n_obs × n_vars = 4518 × 10687 with slots:
feature_id, feature_namecountsdataset_description, dataset_id, dataset_name, dataset_organism, dataset_reference, dataset_summary, dataset_url| Name | Description | Type | Data type | Size |
|---|---|---|---|---|
| var | ||||
feature_
|
Unique identifier for the feature, usually a ENSEMBL gene id. |
vector
|
object
|
10687 |
feature_
|
A human-readable name for the feature, usually a gene symbol. |
vector
|
object
|
10687 |
| layers | ||||
counts
|
Raw counts |
sparsematrix
|
float32
|
4518 × 10687 |
| uns | ||||
dataset_
|
Long description of the dataset. |
atomic
|
str
|
1 |
dataset_
|
A unique identifier for the dataset. This is different from the obs.dataset_id field, which is the identifier for the dataset from which the cell data is derived.
|
atomic
|
str
|
1 |
dataset_
|
A human-readable name for the dataset. |
atomic
|
str
|
1 |
dataset_
|
The organism of the sample in the dataset. |
atomic
|
str
|
1 |
dataset_
|
Bibtex reference of the paper in which the dataset was published. |
atomic
|
str
|
1 |
dataset_
|
Short description of the dataset. |
atomic
|
str
|
1 |
dataset_
|
Link to the original source of the dataset. |
atomic
|
str
|
1 |
dataset.layers['counts']In R: dataset$layers[["counts"]]
Type: sparsematrix, data type: float32, shape: 4518 × 10687
Raw counts
dataset.uns['dataset_description']In R: dataset$uns[["dataset_description"]]
Type: atomic, data type: str, shape: 1
Long description of the dataset.
dataset.uns['dataset_id']In R: dataset$uns[["dataset_id"]]
Type: atomic, data type: str, shape: 1
A unique identifier for the dataset. This is different from the obs.dataset_id field, which is the identifier for the dataset from which the cell data is derived.
dataset.uns['dataset_name']In R: dataset$uns[["dataset_name"]]
Type: atomic, data type: str, shape: 1
A human-readable name for the dataset.
dataset.uns['dataset_organism']In R: dataset$uns[["dataset_organism"]]
Type: atomic, data type: str, shape: 1
The organism of the sample in the dataset.
dataset.uns['dataset_reference']In R: dataset$uns[["dataset_reference"]]
Type: atomic, data type: str, shape: 1
Bibtex reference of the paper in which the dataset was published.
dataset.uns['dataset_summary']In R: dataset$uns[["dataset_summary"]]
Type: atomic, data type: str, shape: 1
Short description of the dataset.
dataset.uns['dataset_url']In R: dataset$uns[["dataset_url"]]
Type: atomic, data type: str, shape: 1
Link to the original source of the dataset.
dataset.var['feature_id']In R: dataset$var[["feature_id"]]
Type: vector, data type: object, shape: 10687
Unique identifier for the feature, usually a ENSEMBL gene id.
dataset.var['feature_name']In R: dataset$var[["feature_name"]]
Type: vector, data type: object, shape: 10687
A human-readable name for the feature, usually a gene symbol.